Le diagnostic de précision au cœur de notre ADN
ID SOLUTIONS
Fabricant français de solutions pour la recherche et le diagnostic moléculaire
ID SOLUTIONS fournit des outils innovants, fiables et standardisés pour la recherche et le développement du diagnostic en oncologie et infectiologie.
De l’extraction d’ADN à la détection par qPCR ou dPCR, nous proposons des solutions complètes qui répondent aux besoins spécifiques des différents acteurs de la recherche et du diagnostic.

Oncologie
Pionnier de la biopsie liquide et de la digital PCR en oncologie
Nous développons nos solutions diagnostiques avec la technologie de dPCR pour répondre aux enjeux de la détection d’ADN tumoral purifié à partir de biopsie liquide (plasma) et solide (échantillons de tissus inclus dans la paraffine (FFPE)) et répondre aux exigences de l’oncologie clinique.
Nous mettons au point des solutions flexibles au service de la recherche en oncologie avec un haut niveau de sensibilité analytique.
Infectiologie
Notre agilité en réponse aux challenges des maladies infectieuses
Le savoir-faire d’ID SOLUTIONS en infectiologie repose sur des outils ergonomiques de pointe, garantissant rapidité, fiabilité, et haute performance dans le dépistage et la prévention des maladies infectieuses.
Nous avons la capacité de proposer des solutions innovantes pour faire face aux situations de crise, en permettant une détection précoce et précise des agents pathogènes.
Nos engagements
La science au service du client

Éthique

Qualité

Satisfaction du client
Actualités
Ne loupez rien des dernières actualités d’ID SOLUTIONS

Recrutement Comptable Général H/F
🔍📊 Opportunité d’emploi : Comptable H/F Qui sommes-nous ? Rejoignez une
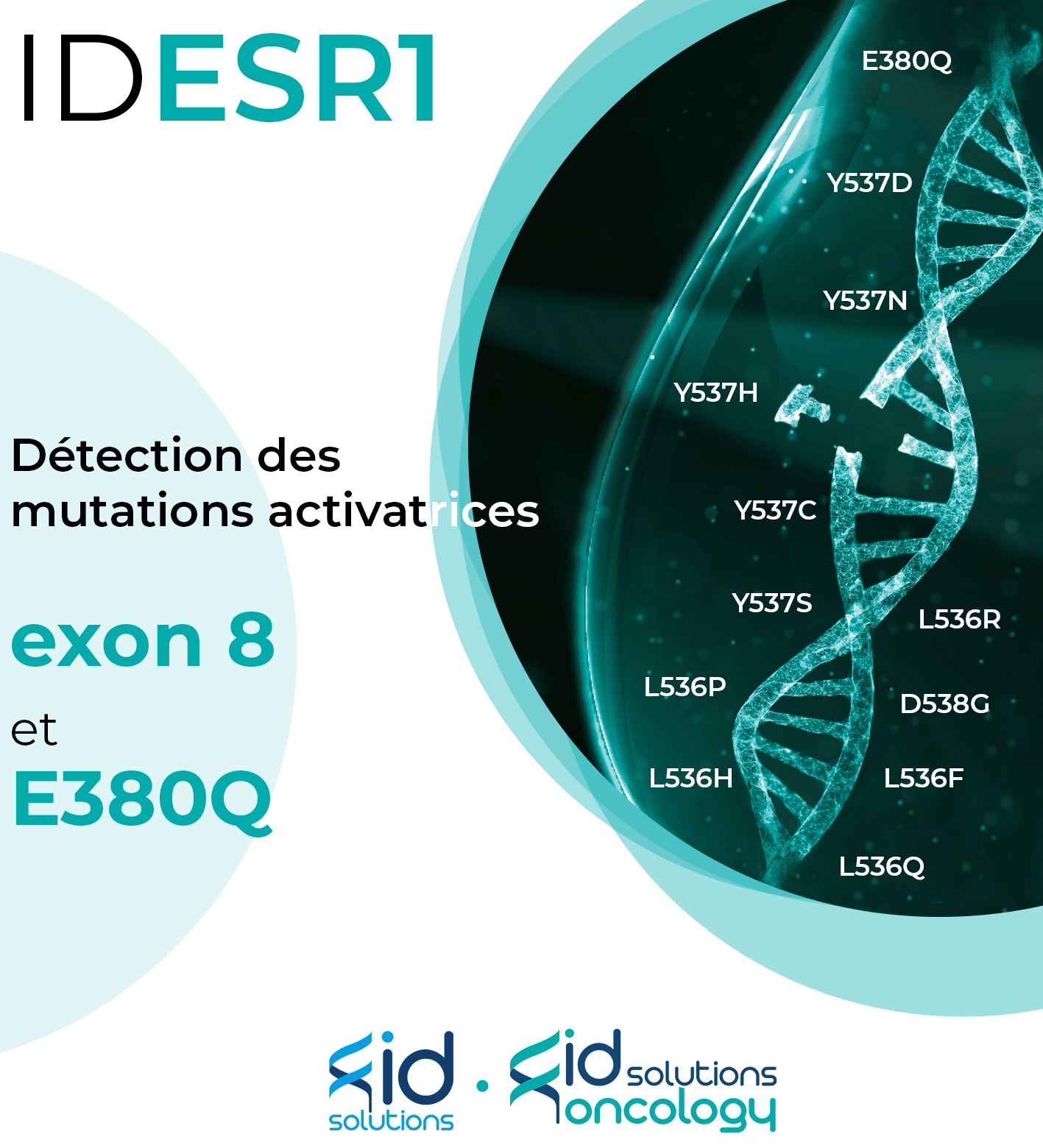
Lancement de notre nouveau kit : IDESR1
ID SOLUTIONS propose des solutions innovantes en biopsie liquide et

On recrute !
⚠ Postes à pourvoir dès que possible ⚠ Vous souhaitez intégrer